Matlab Tools for Biomedical Signal Analysis
Multiscale Entropy
Information Categorization Method
Hilbert-Huang Transform
Physionet toolbox
Multiscale Entropy
Information Categorization Method
Hilbert-Huang Transform
Physionet toolbox
Complexity Analysis of Functional Brain Activity
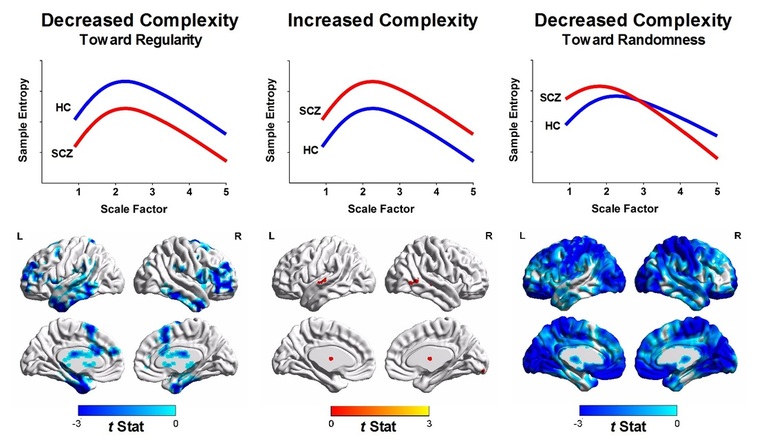
Multiscale entropy (MSE) analysis of resting-state functional magnetic resonance imaging signals (or blood oxygen level dependent BOLD signals). Quantification of complexity changes in patients with schizophrenia (SCZ) relative to healthy controls (HC) according to MSE profiles. (Left) Brain regions with the reduced complexity of BOLD signals toward regularity (i.e., decreased sample entropy over all scale factors in SCZ relative to HC). (Middle) Brain regions with increased complexity of BOLD signals (i.e., increased sample entropy over all scale factors in SCZ relative to HC). Color shown in brain mapping represents raw t-statistics of difference of MSE profile between SCZ and HC. No significant brain cluster was observed in this type of complexity change after correction for multiple comparisons. (Right) Brain regions with reduced complexity toward uncorrelated randomness (i.e., increased sample entropy in short time scales followed by decay of sample entropy as the scale factor increases).
| msentropy.zip | |
| File Size: | 1 kb |
| File Type: | zip |
Download Matlab toolbox for calculating multiscale entropy in EEG and neuroimaging time series data
Installation: unzip all files into a directory (e.g., \msentropy\). Set up the path link in Matlab so the entropy function can be accessed by Matlab console.
Usage: msentropy .m
e = msentropy(input,m,r,factor);
Input: one dimenstion time series vector
m: pattern length or match point(s)
r: similarity parameter
factor: coarsing graining scale factor
Output of msentropy will be one column vector containing sample entropy value for each scale factor.
This multiscale entropy toolbox includes the matlab code of sample entropy which is also available at http://people.ece.cornell.edu/land/PROJECTS/Complexity/sampenc.m
Installation: unzip all files into a directory (e.g., \msentropy\). Set up the path link in Matlab so the entropy function can be accessed by Matlab console.
Usage: msentropy .m
e = msentropy(input,m,r,factor);
Input: one dimenstion time series vector
m: pattern length or match point(s)
r: similarity parameter
factor: coarsing graining scale factor
Output of msentropy will be one column vector containing sample entropy value for each scale factor.
This multiscale entropy toolbox includes the matlab code of sample entropy which is also available at http://people.ece.cornell.edu/land/PROJECTS/Complexity/sampenc.m
Information Categorization (Information-based Similarity Index)
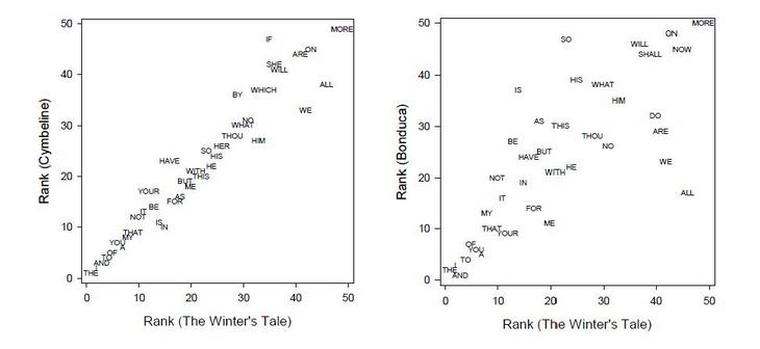
A Matlab toolbox is publicly available at Matlab Central.
The Information-based Similarity (IBS) method was developed to effectively categorize symbolic sequences according to their information content. The method has been fully described and validated, with applications to heart rate time series, literary authorship disputes, and genetic sequences.
This toolbox provides an array of MATLAB functions for quantifying the distance (or dis-similarity) between a set of symbolic sequences, and for displaying the results in graphical form such as dendrogram. The type of symbolic sequences can be binary sequences mapping from a time series, written texts of any given language, genetic sequences, or any symbolic sequences.
In addition to Matlab toolbox, a publicly available software for measuring distance among interbeat interval time series is on Physionet.
PhysioToolkit from PhysioNet

.
PhysioToolkit is a large and growing library of software for physiologic signal processing and analysis, detection of physiologically significant events using both classical techniques and novel methods based on statistical physics and nonlinear dynamics, interactive display and characterization of signals, creation of new databases, simulation of physiologic and other signals, quantitative evaluation and comparison of analysis methods, and analysis of nonequilibrium and nonstationary processes. A unifying theme of the research projects that contribute software to PhysioToolkit is the extraction of "hidden'' information from biomedical signals, information that may have diagnostic or prognostic value in medicine, or explanatory or predictive power in basic research. All PhysioToolkit software is available in source form under the GNU General Public License (GPL).
PhysioToolkit is a large and growing library of software for physiologic signal processing and analysis, detection of physiologically significant events using both classical techniques and novel methods based on statistical physics and nonlinear dynamics, interactive display and characterization of signals, creation of new databases, simulation of physiologic and other signals, quantitative evaluation and comparison of analysis methods, and analysis of nonequilibrium and nonstationary processes. A unifying theme of the research projects that contribute software to PhysioToolkit is the extraction of "hidden'' information from biomedical signals, information that may have diagnostic or prognostic value in medicine, or explanatory or predictive power in basic research. All PhysioToolkit software is available in source form under the GNU General Public License (GPL).
Hilbert-Huang Transform from Research Center for Adaptive Data Analysis
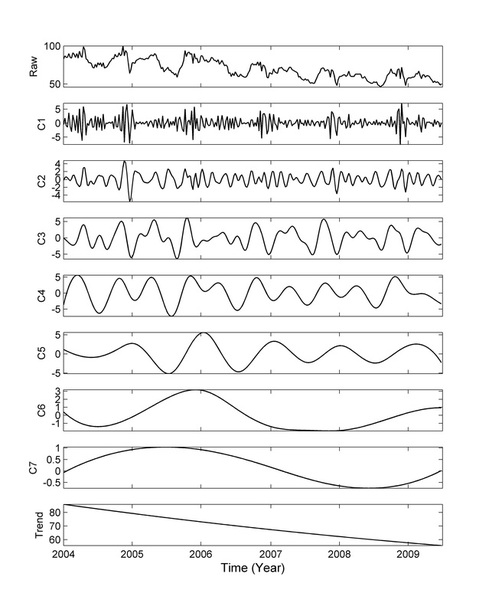
Hilbert-Huang Transform (HHT) is a versatile method developed by Prof. Norden Huang and is used to analyze intricate data from nonlinear and nonstationary processes, including natural and man-made systems. The HHT was selected by NASA for the prestigious Space Act Award and cited as "one of the most important discoveries in the field of applied mathematics in NASA history."
Your comments and suggestions are welcome.
Please send your comments by email to Dr. Albert C. Yang (Laboratory of Precision Psychiatry).
Location: No. 155, Section 2, Linong Street, Beitou District, Taipei, Taiwan
Please send your comments by email to Dr. Albert C. Yang (Laboratory of Precision Psychiatry).
Location: No. 155, Section 2, Linong Street, Beitou District, Taipei, Taiwan


